


 النبات
النبات
 الحيوان
الحيوان
 الأحياء المجهرية
الأحياء المجهرية
 علم الأمراض
علم الأمراض
 التقانة الإحيائية
التقانة الإحيائية
 التقنية الحيوية المكروبية
التقنية الحيوية المكروبية
 التقنية الحياتية النانوية
التقنية الحياتية النانوية
 علم الأجنة
علم الأجنة
 الأحياء الجزيئي
الأحياء الجزيئي
 علم وظائف الأعضاء
علم وظائف الأعضاء
 الغدد
الغدد
 المضادات الحيوية
المضادات الحيوية|
Read More
Date: 19-5-2016
Date: 10-6-2021
Date: 1-11-2015
|
Nucleosomes Are Covalently Modified
KEY CONCEPTS
- Histones are modified by methylation, acetylation, phosphorylation, ubiquitylation, sumoylation, ADPribosylation, and other modifications.
-Combinations of specific histone modifications help to define the function of local regions of chromatin; this is known as the histone code hypothesis.
-The bromodomain is found in a variety of proteins that interact with chromatin; it is used to recognize acetylated sites on histones.
- Several protein motifs recognize methyl lysines, such as chromodomains, PHD domains, and Tudor domains.
All of the histones are subject to numerous covalent modifications, most of which occur in the histone tails. Researchers can modify all of the histones at numerous sites by methylation, acetylation, or phosphorylation, as shown schematically in FIGURE 1. Even though these modifications are relatively small, other, more dramatic modifications occur, as well, such as mono-ubiquitylation, sumoylation, and ADP-ribosylation. Although different histone modifications have known roles in replication, chromatin assembly, transcription, splicing, and DNA repair, researchers have yet to characterize functions of a number of specific modifications.

FIGURE 1 The histone tails can be acetylated, methylated, phosphorylated, and ubiquitylated at numerous sites. Not all possible modifications are shown.
Data from: The Scientist 17 (2003):p. 27.
Lysines in the histone tails are the most common targets of modification. Acetylation, methylation, ubiquitylation, and sumoylation all occur on the free epsilon (ε) amino group of lysine.
As shown in FIGURE 2, acetylation neutralizes the positive charge that resides on the NH3 form of the ε-amino group. In contrast, lysine methylation retains the positive charge, and lysine can be mono-, di-, or trimethylated. Arginine can be mono- or dimethylated. Phosphorylation occurs on the hydroxyl group of serine and threonine. This introduces a negative charge in the form of the phosphate group.

FIGURE 2 The positive charge on lysine is neutralized upon acetylation, whereas methylated lysine and arginine retain their positive charges. Lysine can be mono-, di-, or triacetylated, whereas arginine can be mono- or diacetylated. Serine or threonine phosphorylation results in a negative charge.
All of these modifications are reversible, and a given modification might exist only transiently, or can be maintained stably through multiple cell divisions. Some modifications change the charge of the protein molecule, and, as a result, they are potentially able to change the functional properties of the octamers. For example, extensive lysine acetylation reduces the overall positive charge of the tails, leading to release of the tails from interactions with DNA on their own or other nucleosomes. Modification of histones is associated with structural changes that occur in chromatin at replication and transcription, and specific modifications also facilitate DNA repair. Modifications at specific positions on specific histones can define different functional states of chromatin. Newly synthesized core histones carry specific patterns of acetylation that are removed after the histones are assembled into chromatin, as shown in FIGURE 3. Other modifications are dynamically added and removed to regulate transcription, replication, repair, and chromosome condensation. These other modifications are usually added and removed from histones that are incorporated into chromatin, as depicted for acetylation in FIGURE 4.

FIGURE 3 Acetylation during replication occurs on specific sites on histones before they are incorporated into nucleosomes.
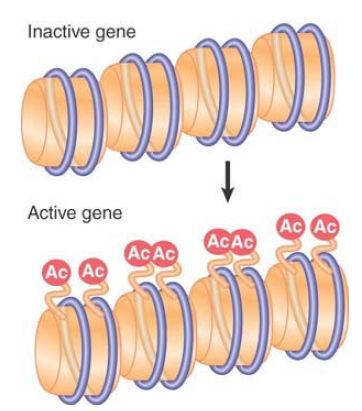
FIGURE 4 Acetylation associated with gene activation occurs by directly modifying specific sites on histones that are already incorporated into nucleosomes.
The specificity of the modifications is controlled by the fact that many of the modifying enzymes have individual target sites in specific histones. TABLE .1 summarizes the effects of some of the modifications that occur on histones H3 and H4. Many modified sites are subject to only a single type of modification in vivo, but others can be subject to alternative modification states (such as lysine 9 of histone H3, which is acetylated or methylated under different conditions). In some cases, modification of one site might activate or inhibit modification of another site. The idea that combinations of signals can be used to define chromatin function led to the idea of a histone code. Although the use of the word “code” has been controversial, this key hypothesis proposes that the collective impact of multiple modifications at particular sites defines the function of a chromatin domain. These modifications are not restricted to a single histone; the functional state of a region of chromatin is derived from all the modifications within a nucleosome or set of nucleosomes. Some modifications of particular histone residues can also prevent or promote other specific histone modification events (or even modification of nonhistone proteins); these “cross-talk” pathways add another level of complexity to signaling through chromatin.
TABLE .1 Most modified sites in histones have a single, specific type of modification, but some sites can have more than one type of modification. Individual functions can be associated with some of the modifications.
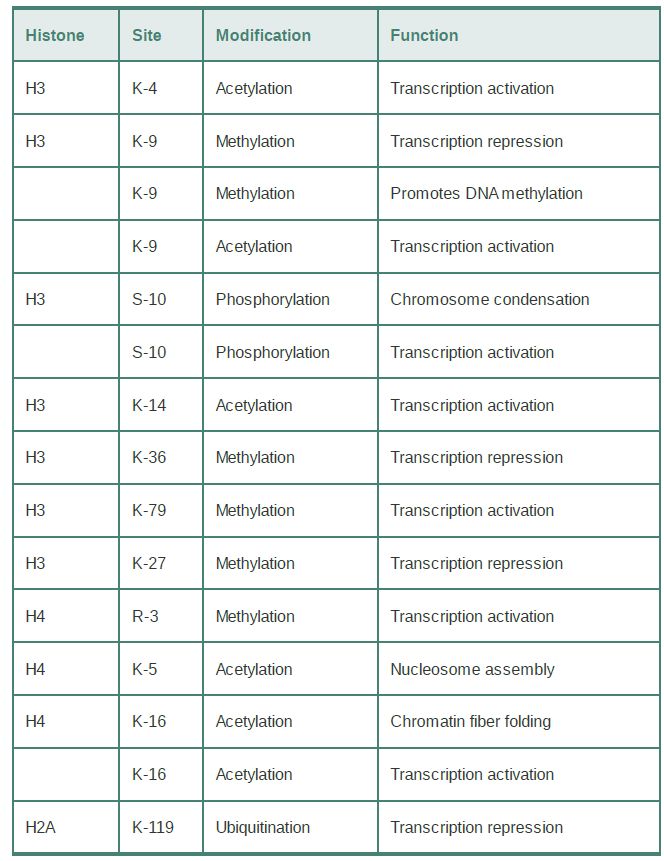
Whereas some histone modifications can directly alter the structure of chromatin, a major function of histone modification lies in the creation of binding sites for nonhistone proteins that change the properties of chromatin. In recent years, a number of protein domains have been identified that bind to specifically modified histone tails. A few examples are provided here.
The bromodomain is found in a variety of proteins that interact with chromatin. Bromodomains recognize acetylated lysine, and different bromodomain-containing proteins recognize different
acetylated targets. The bromodomain itself recognizes only a very short sequence of four amino acids, including the acetylated lysine, so specificity for target recognition must depend on interactions involving other regions. FIGURE 5 shows the structure of a bromodomain bound to its acetylated lysine target. The bromodomain is found in a range of proteins that interact with chromatin, including components of the transcription apparatus and some of the enzymes that remodel or modify histones .
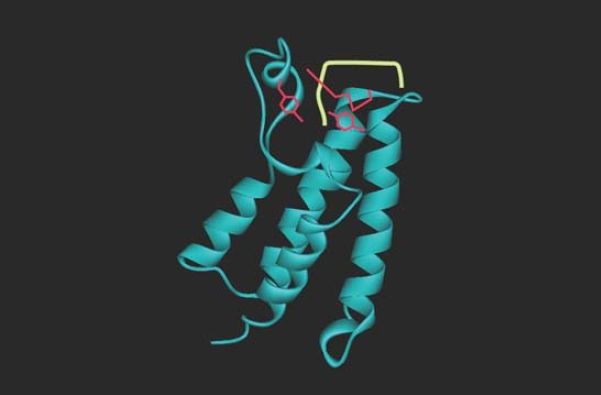
FIGURE 5 Bromodomains are protein motifs that bind acetyllysines. The bromodomain fold consists of a cluster of four α-helices with an acetyl-lysine binding pocket at one end. This figure shows the bromodomain of yeast Gcn5 bound to an H4K16ac peptide.
Data from: Owen, D. J., et al. 2000. “Structure from Protein Data Bank 1E6I.” EMBO J
19:6141–6149.
Methylated lysines (and arginines) are recognized by a number of different domains, which not only can recognize specific modified sites but also can distinguish between mono-, di-, or trimethylated lysines. The chromodomain is a common protein motif of 60 amino acids present in a number of chromatin-associated proteins. Researchers have identified a number of other methyl-lysine binding domains, as shown in FIGURE 6, such as the plant homeodomain (PHD) and the Tudor domain; the number of different motifs designed to recognize particular methylated sites emphasizes the importance and complexity of histone modifications.


FIGURE 6 Numerous protein motifs recognize methylated lysines. (a) The chromodomain of HP1 binds trimethylated K9 of histone H3. (b) The Tudor domain of JMJD2A binds trimethylated K4 of histone H3. Chromodomains and Tudor domains are members of the “royal superfamily,” which bind their targets via a partial β-barrel structure. (c) The PHD finger of BPTF also binds trimethylated K4 of histone H3, using a structure related to DNAbinding zinc finger domains.
(a) Data from: Jacobs, S. A., and Khorasanizadeh, S. 2002. “Structure from Protein Data
Bank 1KNE.” Science 295:2080–2083.
(b) Data from: Y. Huang, et al. 2006. “Structure from Protein Data Bank 2GFA.” Science
12:748–751.
(c) Photo courtesy of Sean D. Taverna, the Johns Hopkins University School of Medicine,
and Haitao Li, Memorial Sloan-Kettering Cancer Center. Additional information at: Taverna,
S. D., et al., Nat Struct Mol Biol 14:1025–1040.
The idea that combinations of modifications are critical, as proposed in the histone code hypothesis, has been reinforced by discoveries of proteins or complexes that can recognize multiple sites of modification simultaneously. For example, some proteins have tandem bromodomains or chromodomains with particular spacing, which can promote binding to histones that are acetylated or methylated at two specific sites. There are also cases in which modification at one site can prevent a protein from recognizing its target modification at another site. It is clear that the effects of a single modification might not always be predictable, and the context of other modifications must be accounted for in order to assign a function to a region of chromatin.

|
|
|
|
تفوقت في الاختبار على الجميع.. فاكهة "خارقة" في عالم التغذية
|
|
|
|
|
|
|
أمين عام أوبك: النفط الخام والغاز الطبيعي "هبة من الله"
|
|
|
|
|
|
|
قسم شؤون المعارف ينظم دورة عن آليات عمل الفهارس الفنية للموسوعات والكتب لملاكاته
|
|
|