Members of a Gene Family Have a Common Organization
 المؤلف:
JOCELYN E. KREBS, ELLIOTT S. GOLDSTEIN and STEPHEN T. KILPATRICK
المؤلف:
JOCELYN E. KREBS, ELLIOTT S. GOLDSTEIN and STEPHEN T. KILPATRICK
 المصدر:
LEWIN’S GENES XII
المصدر:
LEWIN’S GENES XII
 الجزء والصفحة:
الجزء والصفحة:
 9-3-2021
9-3-2021
 2643
2643
Members of a Gene Family Have a Common Organization
KEY CONCEPTS
- A set of homologous genes should share common features that preceded their evolutionary separation.
- All globin genes have a common form of organization with three exons and two introns, suggesting that they are descended from a single ancestral gene.
- Intron positions in the actin gene family are highly variable, which suggests that introns do not separate functional domains.
Many genes in a multicellular eukaryotic genome are related to others in the same genome, either in series (nonallelic) or in parallel (allelic). A gene family is defined as a group of genes that encode related or identical products as a result of gene duplication events. After the first duplication event, the two copies are identical, but then they diverge as different mutations accumulate in them. Further duplications and divergences extend the family. The globin genes are an example of a family that can be divided into two subfamilies (α globin and β globin), but all of its members have the same basic structure and function . In some cases, we can find genes that are more distantly related but that still can be recognized as having common ancestry. Such a group of gene families is called a superfamily.
A fascinating case of evolutionary conservation is presented by the α and β globins and two other proteins related to them. Myoglobin is a monomeric oxygen-binding protein in animals. Its amino acid sequence suggests a common (though ancient) origin with α and β globins. Leghemoglobins are oxygen-binding proteins present in legume plants; like myoglobin, they are monomeric and share a common origin with the other heme-binding proteins. Together, the
globins, myoglobins, and leghemoglobins make up the globin superfamily—a set of gene families all descended from an ancient common ancestor.
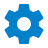
Both α- and β-globin genes have three exons and two introns in conserved positions . The central exon represents the heme-binding domain of the globin chain. There is a single myoglobin gene in the human genome and its structure is essentially the same as that of the globin genes. The conserved three-exon structure therefore predates the common ancestor of the myoglobin and globin genes.
Leghemoglobin genes contain three introns, the first and last of which are homologous to the two introns in the globin genes. This remarkable similarity suggests an exceedingly ancient origin for the interrupted structure of heme-binding proteins, as illustrated in FIGURE 1. The central intron of leghemoglobin separates two exons that together encode the sequence corresponding to the single central exon in globin; the functional heme-binding domain is split into two by an intron. Could the central exon of the globin gene have been derived by a fusion of two central exons in the ancestral gene? Or, is the single central exon the ancestral form? In this case, an intron must have been inserted into it early in plant evolution.
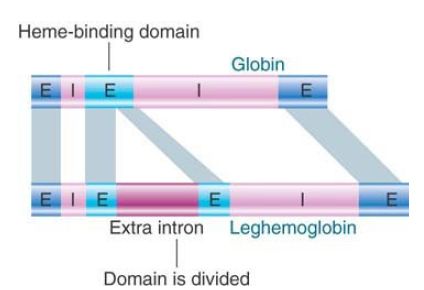
FIGURE 1. The exon structure of globin genes corresponds to protein function, but leghemoglobin has an extra intron in the central domain.
Orthologous genes, or orthologs, are genes that are homologous (homologs) due to speciation; in other words, they are related genes in different species. Comparison of orthologs that differ in structure might provide information about their evolution. An example is insulin. Mammals and birds have only one gene for insulin, except for rodents, which have two. FIGURE 2. illustrates the structures of these genes.

FIGURE 2. The rat insulin gene with one intron evolved by loss of an intron from an ancestor with two introns.
We use the principle of parsimony in comparing the organization of orthologous genes by assuming that a common feature predates the evolutionary separation of the two species. In chickens, the single insulin gene has two introns; one of the two homologous rat genes has the same structure. The common structure implies that the ancestral insulin gene had two introns. However, because the second rat gene has only one intron, it must have evolved by a gene duplication in rodents that was followed by the precise removal of one intron from one of the homologs.
The organizations of some orthologs show extensive discrepancies between species. In these cases, there must have been extensive deletion or insertion of introns during evolution. A well characterized case is that of the actin genes. The common features of actin genes are an untranslated leader of fewer than 100 bases, a coding region of about 1,200 bases, and a trailer of about 200 bases. Most actin genes have introns, and their positions can be aligned with regard to the coding sequence (except for a single intron sometimes found in the leader).
FIGURE 3. shows that almost every actin gene is different in its pattern of intron positions. Among all the genes being compared, introns occur at 19 different sites. However, the range of intron number per gene is zero to six. How did this situation arise? If we suppose that the ancestral actin gene had introns, and that all current actin genes are related to it by loss of introns, different introns have been lost in each evolutionary branch. Probably some introns have been lost entirely, so the ancestral gene could well have had 20 introns or more. The alternative is to suppose that a process of intron insertion continued independently in the different lineages.

FIGURE 3. Actin genes vary widely in their organization. The sites of introns are indicated by dark boxes. The bar at the top summarizes all the intron positions among the different orthologs.
Whether introns were present in actin genes early or late, there appears to have been no consistent influence from actin protein domains or subdomains as to where introns should be located. On the other hand, when exons are under negative selection (resulting in homology conservation), in-series recombination between members of an expanding gene family (that could cause a contraction in family size) would be decreased by intron diversification (resulting in loss of some homology), and introns would come to reside where this could best be achieved.
Alleles would have similar exons and introns, so in-parallel interallelic recombination (as in meiosis) would be unimpaired until speciation occurred—a process that could be accompanied by intron relocations. The relationships between the intron locations among different species could then be used to construct a phylogenetic tree illustrating the evolution of the actin gene.
The relationship between individual exons and functional protein domains is somewhat erratic. In some cases, there is a clear oneto-one relationship; in others, no pattern can be discerned. One possibility is that the removal of introns has fused the previously adjacent exons. This means that the intron must have been precisely removed without changing the integrity of the coding region. An alternative is that some introns arose by insertion into an exon encoding a single domain. Together with the variations that we see in exon placement in cases such as the actin genes, the conclusion is that intron positions can evolve.
The correspondence of at least some exons with protein domains and the presence of related exons in different proteins leave no doubt that the duplication and juxtaposition of exons have played important roles in evolution. It is possible that the number of ancestral exons—from which all proteins have been derived by duplication, variation, and recombination—could be relatively small, perhaps as little as a few thousand. The idea that exons are the building blocks of new genes is consistent with the “introns early” model for the origin of genes encoding proteins .
 الاكثر قراءة في مواضيع عامة في الاحياء الجزيئي
الاكثر قراءة في مواضيع عامة في الاحياء الجزيئي
 اخر الاخبار
اخر الاخبار
اخبار العتبة العباسية المقدسة


